missing translation for 'onlineSavingsMsg'
Learn More
Learn More
Invitrogen™ AKAP10 Polyclonal Antibody


Description
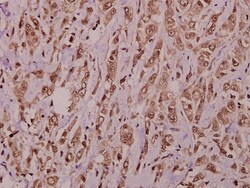
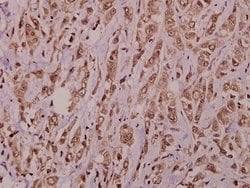
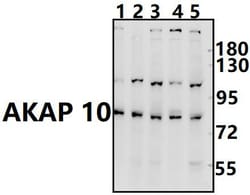
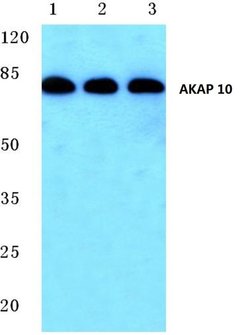
This antibody detects endogenous protein at a molecular weight of 74 kDa. Purity is >95% by SDS-PAGE.
The A-kinase anchor proteins are a group of structurally diverse proteins, which have the common function of binding to the regulatory subunit of protein kinase A and confining the holoenzyme to discrete locations within the cell. This gene encodes a member of the AKAP family. The encoded protein interacts with both the type I and type II regulatory subunits of PKA; therefore, it is a dual-specific AKAP. This protein is highly enriched in mitochondria. It contains RGS domains, in addition to a PKA-RII subunit-binding domain. The mitochondrial localization and the presence of RGS domains may have important implications for the function of this protein in PKA and G protein signal transduction.

Specifications
Specifications
| Antigen | AKAP10 |
| Applications | Immunohistochemistry (Paraffin), Western Blot |
| Classification | Polyclonal |
| Concentration | 1 mg/mL |
| Conjugate | Unconjugated |
| Formulation | PBS with 50% glycerol and 0.02% sodium azide; pH 7.2 |
| Gene | AKAP10 |
| Gene Accession No. | O43572 |
| Gene Alias | 1500031L16Rik; A kinase (PRKA) anchor protein 10; A kinase anchor protein 10; A kinase anchor protein 10, mitochondrial; Akap10; AKAP-10; A-kinase anchor protein 10, mitochondrial; A-kinase anchoring protein 10; B130049N18Rik; D-akap2; D-AKAP-2; Dual specific A kinase anchoring protein 2; dual specificity A kinase-anchoring protein 2; MGC9414; mitochondrial A kinase PPKA anchor protein 10; PRKA10; protein kinase A anchoring protein 10; protein kinase A-anchoring protein 10 |
| Gene Symbols | AKAP10 |
| Show More |
Product Title
By clicking Submit, you acknowledge that you may be contacted by Fisher Scientific in regards to the feedback you have provided in this form. We will not share your information for any other purposes. All contact information provided shall also be maintained in accordance with our Privacy Policy.
Spot an opportunity for improvement?